Chapter 2 Getting started
2.1 System requirements
Osprey requires MATLAB and has been tested on version 2017a and newer (2019a and newer is required for the GUI). The following toolboxes are required for full functionality:
- Optimization
- Statistics and Machine Learning
You can check your current MATLAB version and the available toolboxes with the following command at the MATLAB prompt:
-----------------------------------------------------------------------------------------------------
MATLAB Version: 9.4.0.949201 (R2018a) Update 6
MATLAB License Number: 703789
Operating System: Mac OS X Version: 10.14.6 Build: 18G4032
Java Version: Java 1.8.0_144-b01 with Oracle Corporation Java HotSpot(TM) 64-Bit Server VM mixed mode
-----------------------------------------------------------------------------------------------------
MATLAB Version 9.4 (R2018a)
Curve Fitting Toolbox Version 3.5.7 (R2018a)
GUI Layout Toolbox Version 2.3.4 (R2018b)
Global Optimization Toolbox Version 3.4.4 (R2018a)
Image Processing Toolbox Version 10.2 (R2018a)
Neural Network Toolbox Version 11.1 (R2018a)
Optimization Toolbox Version 8.1 (R2018a)
Parallel Computing Toolbox Version 6.12 (R2018a)
Signal Processing Toolbox Version 8.0 (R2018a)
Statistics and Machine Learning Toolbox Version 11.3 (R2018a)
Symbolic Math Toolbox Version 8.1 (R2018a)
Wavelet Toolbox Version 5.0 (R2018a)
Widgets Toolbox Version 1.3.330 (R2019a)2.2 Installing Osprey and necessary dependencies
2.2.1 Osprey package
Get the latest Osprey code from our GitHub repository:
Clone the repository, or download and extract the contents of the .ZIP file to a directory on your drive.
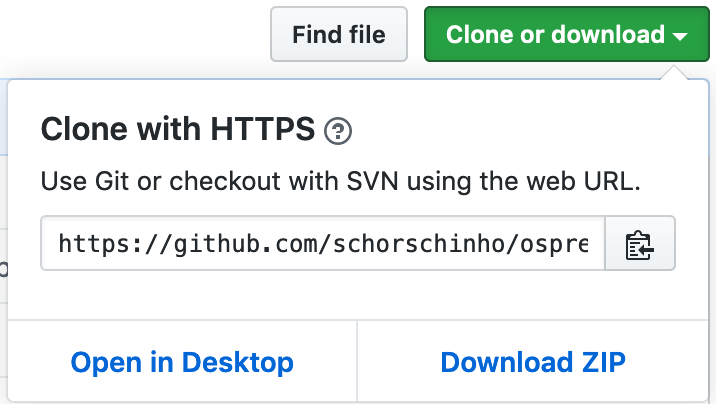
Figure 2.1: Cloning Osprey from GitHub.
Add the entire
ospreydirectory (with subfolders) to your MATLAB path.Make sure to regularly check the repository for updates, as we frequently commit new features, bug fixes, and improved functions.
2.2.2 SPM package
To perform voxel co-registration and tissue segmentation, download SPM12 from the University College London website. You will need to provide the SPM team with some information before you can access the download link.
- Extract the downloaded archive so that the
spm12folder is on the same directory level as theospreyfolder:

Figure 2.2: Make sure that the SPM12 and Osprey folders are in the same directory.
- Add the
spm12folder to your MATLAB path, but without subfolders.
2.2.3 GUI Toolboxes
If you want to use the Osprey Graphical User Interface (GUI), please download the following toolboxes from the MATLAB File Exchange:
GUI Layout Toolbox (David Sampson)
Widget Toolbox (Robin Jackey)
Download both toolboxes in the MATLAB toolbox format (.mltbx). You can double-click to install. MATLAB will automatically add the toolboxes to its path.
2.3 How to organize your raw data
Raw MRS data come in an overwhelming variety of formats, each producing different numbers of files. Osprey does not make a lot of assumptions with regard to your folder structure, since you specify the exact location for each file in a job file prior to each analysis. It is highly recommended, however, that you store different acquisitions in separate folders. Organizing your raw data in a consistent and meaningful way will save you a lot of time and nerves.
To ensure optimal functioning of Osprey, we suggest adapting the folder hierarchy proposed by the BIDS (Brain Imaging Data Structure) initiative:
There are four main levels of the folder hierarchy, these are:
project/
└── subject
└── session
└── acquisitionWith the exception of the top-level
projectfolder, all sub-folders have a specific structure to their name (described below). Here’s an example of how this hierarchy looks:
myProject/
└── sub-01
└── ses-01
└── anatAn example adaptation of this hierarchy for Philips MRS data could look like this:
myProject/
├── sub-01
│ └── ses-01
│ ├── anat
│ │ └── sub-01_T1w.nii
│ └── mrs
│ ├── sub-01_mega-press_act
│ │ ├── sub-01_mega-press_act.sdat
│ │ └── sub-01_mega-press_act.spar
│ ├── sub-01_mega-press_ref
│ │ ├── sub-01_mega-press_ref.sdat
│ │ └── sub-01_mega-press_ref.spar
│ └── sub-01_press-water
│ ├── sub-01_press-water_act.sdat
│ └── sub-01_press-water_act.spar
├── sub-02
│ └── ses-01
│ ├── anat
│ │ └── sub-02_T1w.nii
│ └── mrs
│ ├── sub-02_mega-press_act
│ │ ├── sub-02_mega-press_act.sdat
│ │ └── sub-02_mega-press_act.spar
│ ├── sub-02_mega-press_ref
│ │ ├── sub-02_mega-press_ref.sdat
│ │ └── sub-02_mega-press_ref.spar
│ └── sub-02_press-water
│ ├── sub-02_press-water_act.sdat
│ └── sub-02_press-water_act.spar
...
...While adhering to the BIDS standard is strongly recommended, it is by no means
binding for most file formats. You can, for example, keep Siemens TWIX files
(.dat) all in the same folder, and Osprey will be able to handle them, if
you define their paths in the job file accordingly.
*.IMA, *.DCM) or single-average Siemens RDA format, it is absolutely necessary to keep every scan in a separate folder.